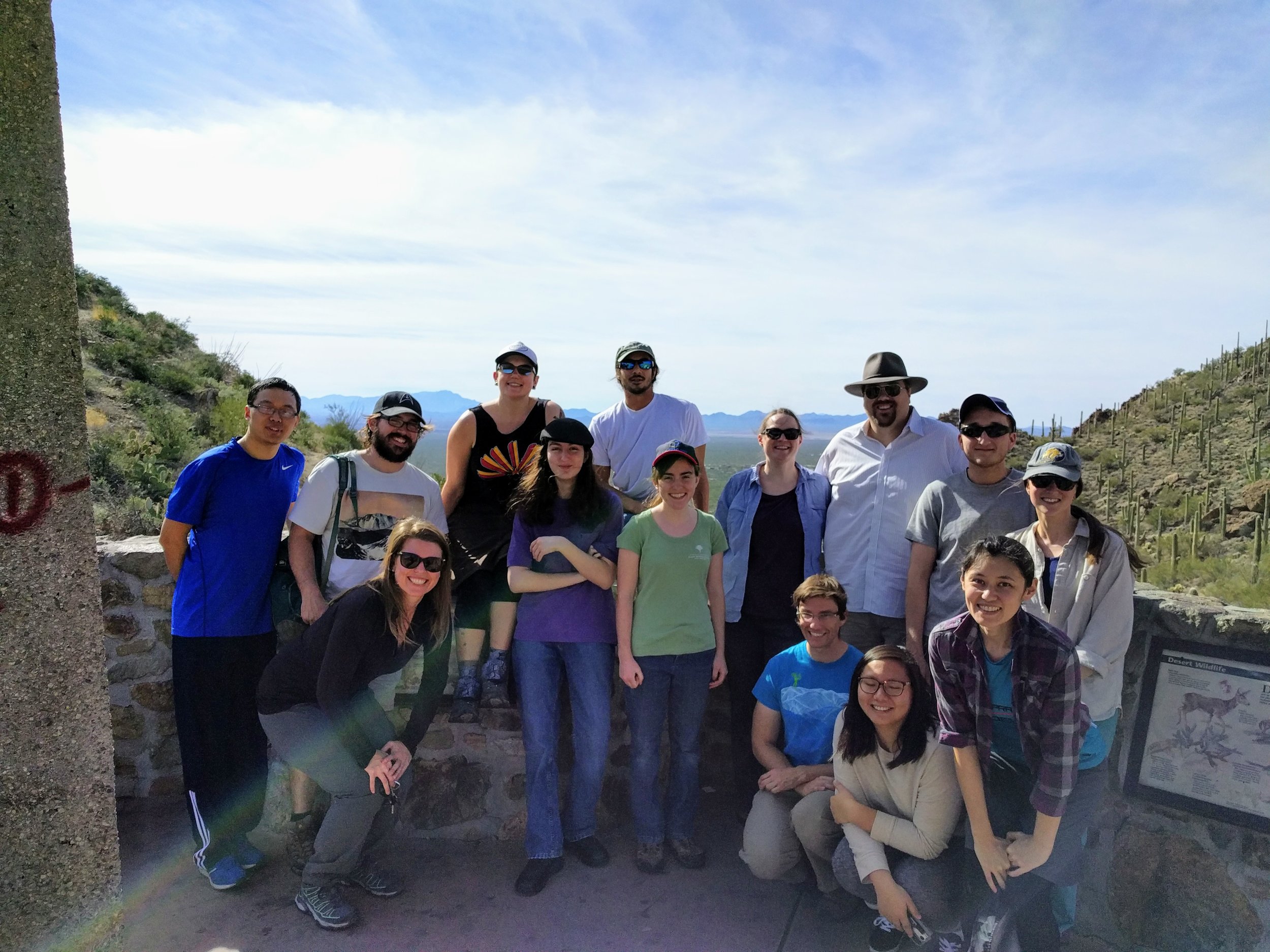
People
Most of the 2025 Barker Lab at Botany 2025 in Palm Springs!
Dr. Michael S. Barker
Professor and Associate Department Head, Ecology & Evolutionary Biology
Professor, BIO5
Professor, Genetics GIDP
Director, Bioinformatics Degree Program
Co-Director, CAMBIUM NRT
msbarker@arizona.edu
Phone: 520–621–2213
Office: BioSciences West 321E
Lab: BioSciences West 321
EEB Department Page
@barkerms.bsky.social on Bluesky
Dr. Tamsen Dunn
NSF PRFB Postdoctoral Fellow, January 2026
Developing computational approaches to study the evolution of polyploid genomes
Geoff Finch
EEB Ph.D. Candidate
Analyzing the population genetic consequences of chromosome number evolution with a focus on the exceptional genus Xanthisma
Seongyeon Kang
EEB Ph.D. Student
Exploring chromosomal evolution in flowering plants
Alex Stewart
EEB PhD Student
NSF GRFP Fellow
Studying polyploid speciation and genome evolution in salamanders and Selaginella
Polly Barker
Lab mascot and guard dog of BSW 321E
Lab Alumni
Postdoctoral Researchers
Dr. Justin Conover (NSF PRFB Postdoctoral Fellow, 2022-2025) → Currently Assistant Member and Principal Investigator at the Donald Danforth Plant Science Center, St. Louis
Dr. Sylvia Kinosian (NSF PRFB Postdoctoral Fellow, 2022-2025) → Currently Assistant Professor and Herbarium Curator at Montana State University
Dr. Jay Goldberg (NSF PRFB Postdoctoral Fellow, 2020-2023) → Currently ASU Presidential Postdoctoral Fellow
Dr. Paul Blischak (NSF Plant Genome Postdoctoral Fellow, 2018-2020) → Currently Data Scientist at Bayer Crop Science
Dr. Brittany Sutherland (NIH PERT Postdoctoral Fellow, 2017-2019) → Currently Assistant Professor at George Mason University
Dr. Hannah Marx (2016-2018) → Currently Director of the L.H. Bailey Herbarium and Assistant Professor at Cornell University
Dr. Xinshuai Qi (2014-2017) → Currently Bioinformatics Consultant at Deloitte
Dr. Emily Sessa (2012-2013) → Currently Patricia K. Holmgren Director of the William and Lynda Steere Herbarium at the New York Botanical Garden
Dr. Pat Edger (NSF Plant Genome Postdoctoral Fellow, 2012-2015) → Currently Associate Professor in the Department of Horticulture at Michigan State University
Dr. Rebecca Rundell (G.G. Simpson Postdoctoral Fellow, 2011-2012) → Currently Associate Professor at SUNY-ESF, Syracuse, NY
Dr. Nils Arrigo (Swiss FNS Postdoctoral Fellow, 2010-2012) → Senior Bioinformatician at Sophia Genetics
Graduate Students
Dr. Michael McKibben (EEB Ph.D., 2019-2025) → Currently postdoctoral fellow at the Donald Danforth Plant Science Center
Qiuyu Jiang (EEB M.S., 2021-2024)
Andrew Vizzerra (EEB M.S., 2020-2022)
Dr. Zheng Li (EEB Ph.D., 2013-2020) → Currently Assistant Professor and Herbarium Curator at Miami University, Oxford, Ohio
Dr. Anthony Baniaga (EEB Ph.D., 2012-2019) → Currently Herbarium Curator at University of California, Los Angeles
Stacey Jorgensen (EEB 2014-2017)
Undergraduates
Sophia Delgado (2024-2025)
Cole Fields (2022-2023)
CeeCee Hill (2022-2023)
Patrick Hsu (2021-2022)
Summer Dow (2020)
Will Nenad (2020-2022)
Yang Yang (2020-2021)
Ellie Asadi (2017-2019)
Tatiana Nzeukou (2018-2019)
Alisa Di Pasqua (2018-2019)
Denae Yutarte (2018-2019)
Kyle Jenkins (2017-2018)
Alex Trostle (2016-2018)
Anshu Goyal (2016-2018)
Alexandra Lei Pond (2016-2018)
Sally Galuska (2016-2017)
Chris Reardon (2014-2017)
Tara Hall (2014-2015)
Thomas Kidder (2014-2017)
Rob Cerasa (2014-2015)
Kassidi Retkowski (2014-2015)
Zainab Sadeq (2014-2015)
Kaitlin Ann Libby (2013-2014)
Mandolina Fitzgibbon (2013)
Kevin Holt (2012-2014)
Rick Russell (2012-2013)
Jolene Nelson (2011-2012)
Rishi Masalia (2011-2012)
Shing Zhan (UBC 2008-2011)
Visiting Scholars
Carlos A. Flores López
Jill Hamilton
George Tiley
Jesse Czekanski-Moir
Hanghui Kong
Rongpei Yu
Elton Assis
Kay Hodgins